295,00 € – 995,00 €
Product details
Synonyms = GC79; LGCR; Transcriptional repressor GATA binding 1; Tricho-rhino-phalangeal syndrome type I protein; Trichorhinophalangeal syndrome I; Trichorhinophalangeal syndrome I homolog; TRPS 1; trpS1; TRPS1 gene; TRPS1_HUMAN; Zinc finger protein GC79; Zinc finger transcription factor TRPS 1 antibody
Antibody type = Rabbit monoclonal / IgG
Clone = MSVA-512R
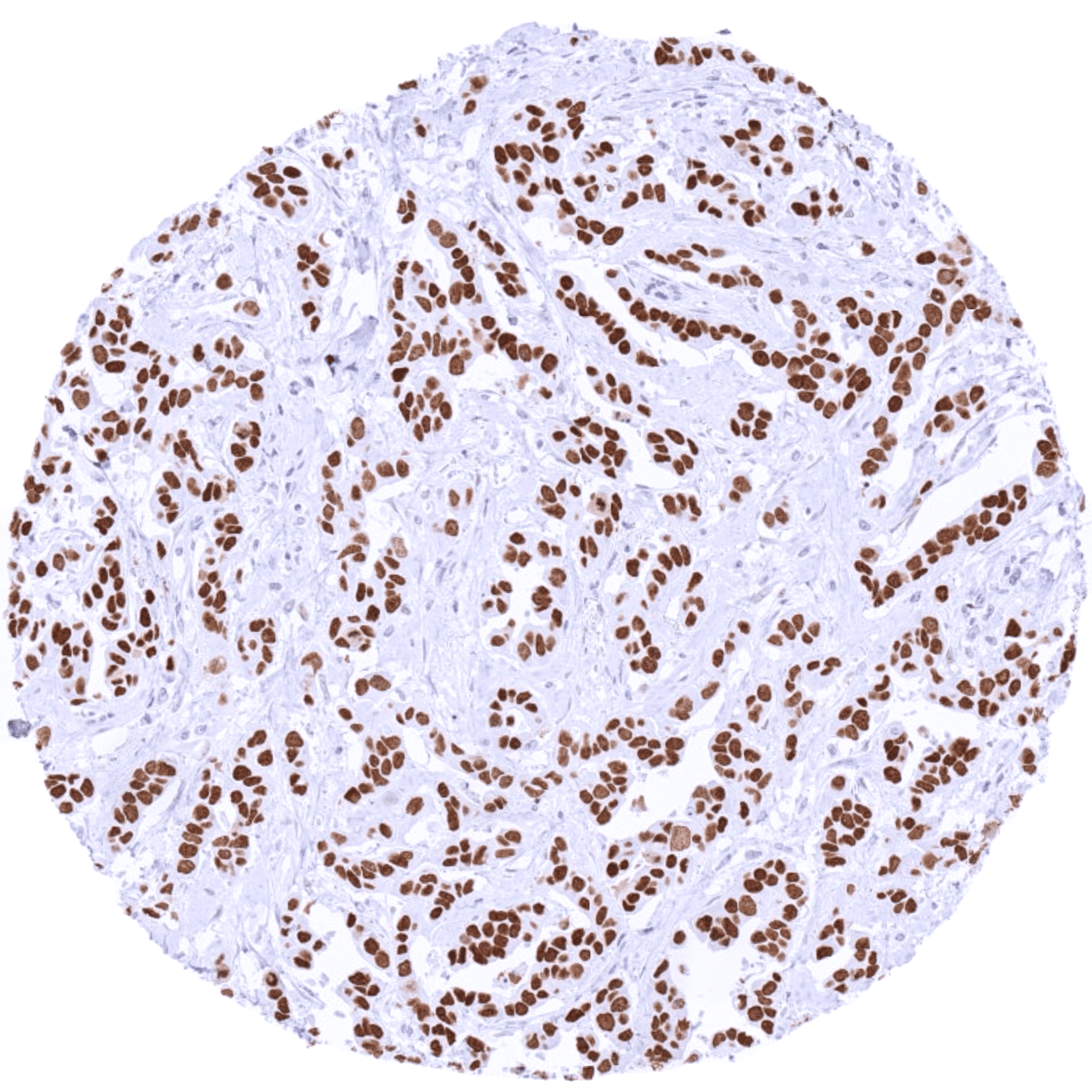
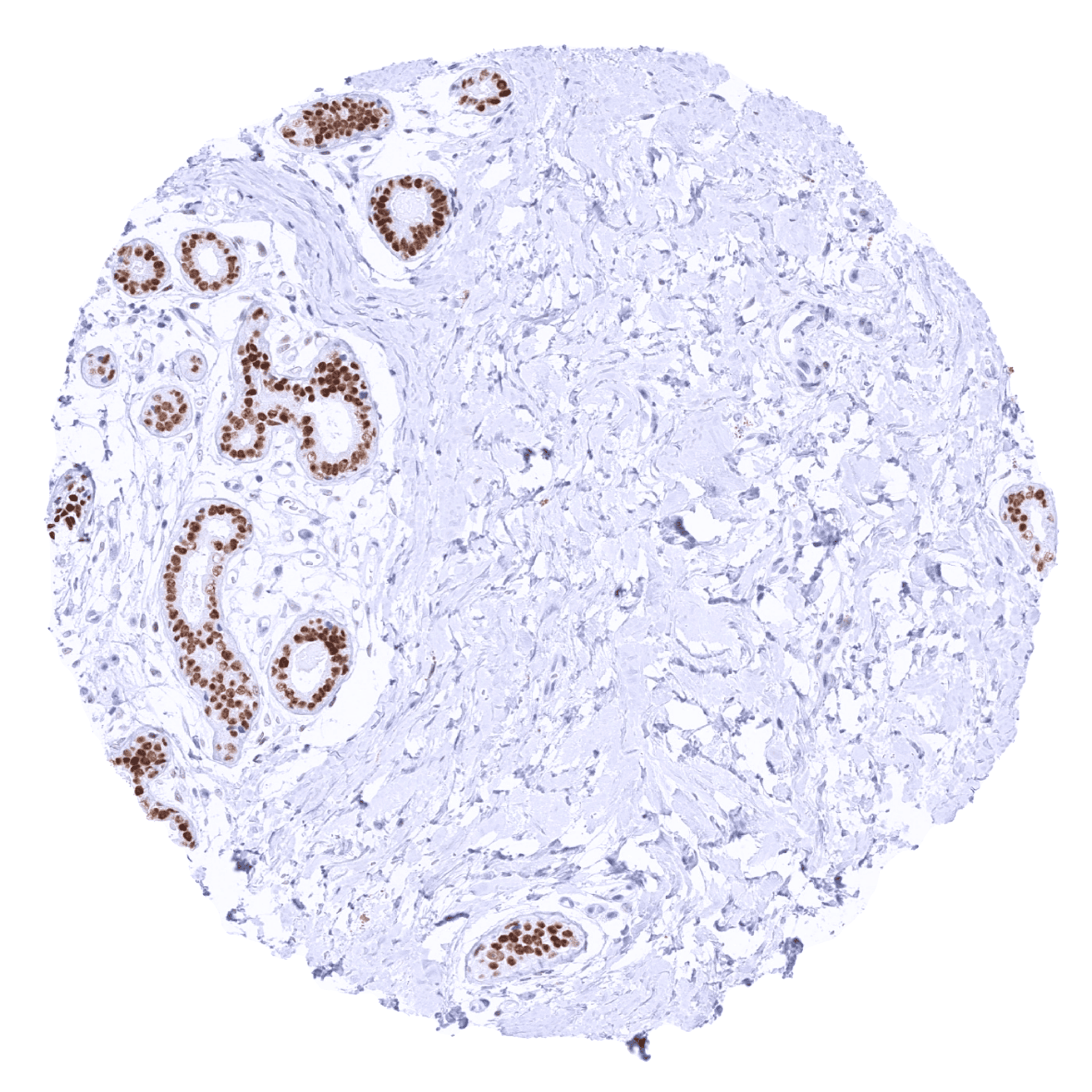
Positive control = Breast: A strong TRPS1 staining should be seen in luminal breast epithelial cells. (For low-level TRPS1 detection: Testis: A weak nuclear TRPS1 staining should be seen in spermatogonia.)
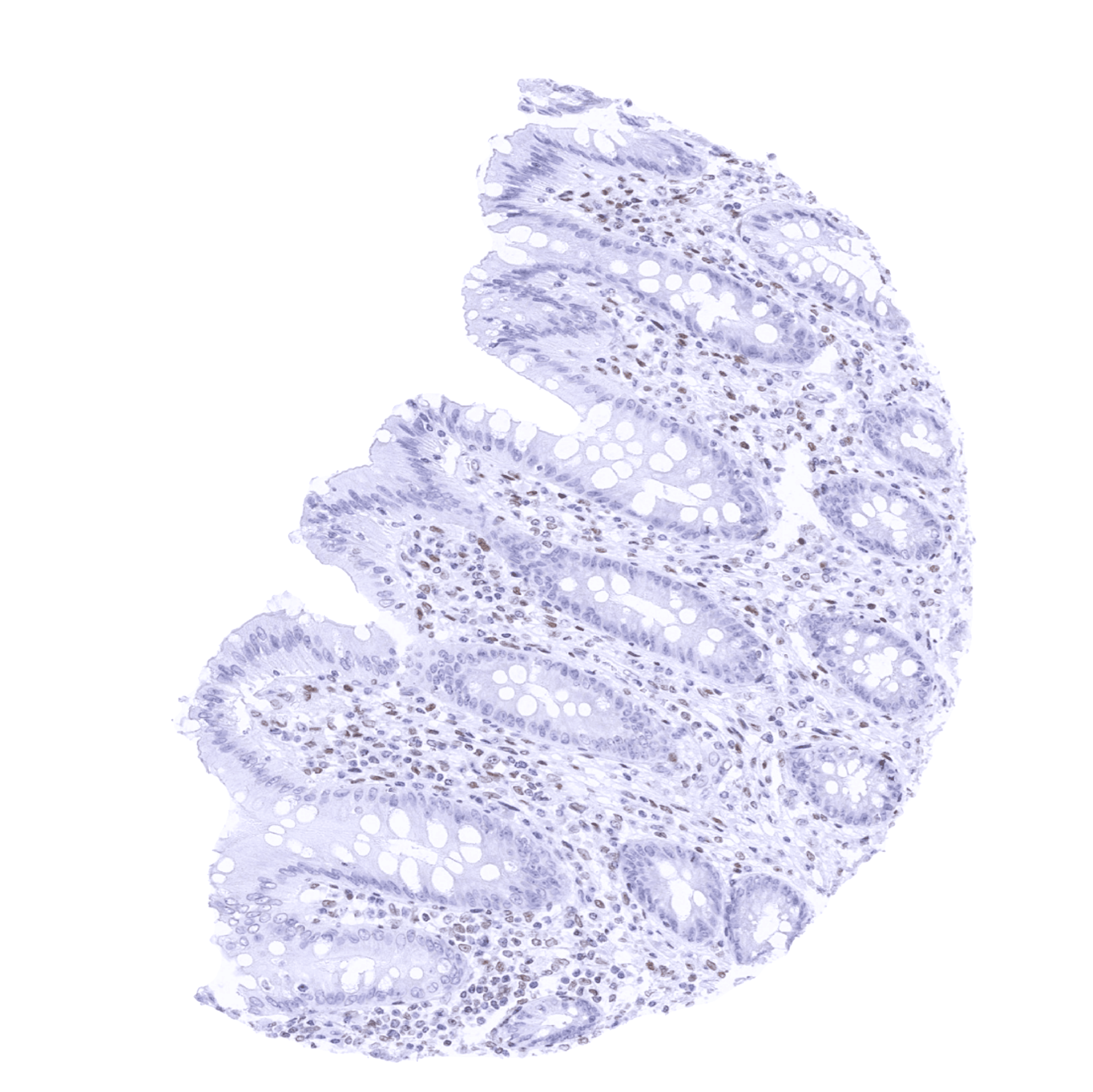
Negative control = Colon: Nuclear TRPS1 staining should be absent in epithelial cells.
Cellular localization = Nucleus
Reactivity = Human
Application = Immunohistochemistry
Dilution = 1:100 – 1:200
Intended Use = Research Use Only
Relevance of Antibody
TRPS1 is a Transcription factor highly expressed in breast epithelial tissues.
Biology Behind
TRPS1 (Transcriptional Repressor GATA Binding 1) is a nuclear transcription factor protein coded by the TRPS1 gene on chromosome 8q23-24. In contrast to other GATA-type transcription factors, TRPS1 mainly acts as a transcriptional repressor. TRPS1 represses GATA-regulated genes by binding to the dynein light chain protein. TRPS1-dynein binding hinders dynein-GATA binding and thus suppresses its transcriptional activity. For example, TRPS1 can directly suppress the expression of Runx1 and Sox9 in cartilage formation. Several hereditary TRPS1 mutations are known to result in craniofacial and skeletal malformations. TRPS1 can also induce expression of genes such as for example FOXA1, a negative regulator of epithelial mesenchymal transition (EMT). It was suggested that TRPS1 has tumor-suppressive activity by preventing EMT. Aberrant expression of TRPS1 occurs in various types of cancers.
Staining Pattern in Normal Tissues
Images describing the TRPS1 staining pattern in normal tissues obtained by the antibody MSVA-512R are shown in our “Normal Tissue Gallery”.
| Brain | Cerebrum | Moderate nuclear TRPS1 staining of glia cells. |
| Cerebellum | Moderate nuclear TRPS1 staining of glia cells. | |
| Endocrine Tissues | Thyroid | Faint nuclear TRPS1 staining of a fraction of epithelial cells. |
| Parathyroid | Weak to moderate nuclear TRPS1 staining of epithelial cells. | |
| Adrenal gland | Negative. | |
| Pituitary gland | Faint nuclear TRPS1 staining of a fraction of epithelial cells. | |
| Respiratory system | Respiratory epithelium | Weak to moderate nuclear TRPS1 staining of a fraction of respiratory epithelial cells and of some glandular cells from bronchus glands. |
| Lung | Negative. | |
| Gastrointestinal Tract | Salivary glands | Weak nuclear TRPS1 staining in glandular (serous and mucinous) and excretory duct cells. |
| Esophagus | Weak to moderate nuclear TRPS1 staining of suprabasal squamous epithelial cells. | |
| Stomach | Epithelial cells are TRPS1 negative. | |
| Duodenum | Epithelial cells and Brunner glands are negative. | |
| Small intestine | Negative. Stroma cells can be positive. | |
| Appendix | Negative. Stroma cells can be positive. | |
| Colon | Negative. Stroma cells can be positive. | |
| Rectum | Negative. Stroma cells can be positive. | |
| Liver | Negative. | |
| Gallbladder | A faint nuclear TRPS1 staining can occur in epithelial cells. | |
| Pancreas | A weak nuclear TRPS1 staining can occur in stroma cells. | |
| Genitourinary | Kidney | Weak nuclear TRPS1 staining in a fraction of cells in tubuli and collecting ducts. |
| Urothelium | Weak nuclear TRPS1 staining in a fraction of urothelial cells. | |
| Male genital | Prostate | Faint to moderate nuclear TRPS1 staining can occur in acinar cells. Probably depending on functional status and perhaps preferentially in atrophic glands. Weak to moderate staining of (muscular) stroma cells. |
| Seminal vesicles | Variable (weak to strong) nuclear TRPS1 staining in muscular stroma cells. | |
| Testis | Weak nuclear TRPS1 staining in spermatogonia. | |
| Epididymis | Very faint nuclear TRPS1 staining in some epithelial cells. | |
| Female genital | Breast | Intense nuclear TRPS1 staining of luminal (but not basal) epithelial cells. |
| Uterus, myometrium | Weak nuclear TRPS1 staining in a fraction of muscular cells. | |
| Uterus, ectocervix | Weak to moderate nuclear TRPS1 staining of suprabasal squamous epithelial cells. | |
| Uterus endocervix | Epithelial cells are usually negative but can be TRPS1 positive (probably in case of “stressed” epithelium”) | |
| Uterus, endometrium | Moderate nuclear TRPS1 staining of epithelial cells. A somewhat weaker staining is seen in stroma cells. | |
| Fallopian Tube | Weak nuclear TRPS1 staining of a fraction of epithelial cells. | |
| Ovary | Weak nuclear TRPS1 staining in a fraction of stroma cells. | |
| Placenta early | Negative. | |
| Placenta mature | Negative. | |
| Amnion | A weak nuclear TRPS1 staining can rarely be seen. | |
| Chorion | Negative. | |
| Skin | Epidermis | Weak nuclear TRPS1 staining can be seen in suprabasal squamous epithelial cells (not in all samples). |
| Sebaceous glands | Moderate nuclear TRPS1 staining of sebaceous gland cells and in epithelial cells of hair follicles. | |
| Muscle/connective tissue | Heart muscle | Moderate nuclear TRPS1 staining of a fraction of stroma cells. |
| Skeletal muscle | Weak nuclear TRPS1 staining of a fraction of stroma cells. | |
| Smooth muscle | Weak to moderate nuclear TRPS1 staining in many organs. | |
| Vessel walls | Usually negative. Some spindle cells in the aortic wall show weak to moderate TRPS1 staining. | |
| Fat | Negative. | |
| Stroma | Weak to moderate nuclear TRPS1 staining can occur in stroma cells (fibroblasts?) | |
| Endothelium | Negative. | |
| Bone marrow/lymphoid tissue | Bone marrow | Negative. |
| Lymph node | Negative. | |
| Spleen | Negative. | |
| Thymus | Negative. | |
| Tonsil | Lymphocytes are negative. | |
| Remarks | Staining intensity is highest in breast epithelial cells but many other cell types do also show staining. |
These findings are largely consistent with the RNA data described in the Human Protein Atlas (Tissue expression TRPS1)
TRPS1 staining by MSVA-512R is most prominent in breast tissue but also occurs – at lower levels – in a very broad range of other tissues.
Positive control = Breast: A strong TRPS1 staining should be seen in luminal breast epithelial cells. (For low-level TRPS1 detection: Testis: A weak nuclear TRPS1 staining should be seen in spermatogonia.)
Negative control = Colon: Nuclear TRPS1 staining should be absent in epithelial cells.
Staining Pattern in Relevant Tumor Types
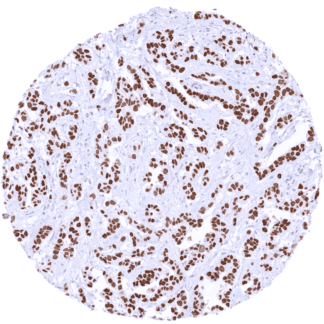
A strong TRPS1 immunostaining preferentially occurs in breast cancer. TRPS1 positivity – usually at a lower level of intensity – also occurs in many other tumor entities derived from the female genital tract and from various other organs. If TRPS1 should be used as a marker for breast cancer, a much higher dilution is recommended than for use for low-level TRPS1 detection.
The TCGA findings on TRPS1 RNA expression in different tumor categories have been summarized in the Human Protein Atlas.
Compatibility of Antibodies
TRPS1 (MSVA-512R) publication summary
Relevant publication: Lennartz et al. “TRPS1 is a highly sensitive marker for breast cancer: A tissue microarray study evaluating more than 19,000 tumors from 152 different tumor entities” Published in Am J Surg Pathol. 2024 Apr 22. doi: 10.1097/PAS.0000000000002213. Epub ahead of print. PMID: 38647255.
A total of 16,818 tumors from 152 different tumor categories were successfully analyzed by using the following protocol: Heat-induced antigen retrieval for 5 minutes in an autoclave at 121°C in pH 9,0 Target Retrieval Solution buffer. MSVA-512R, at a dilution of 1:300 at 37°C for 60 minutes. Visualization of bound antibody by the EnVision Kit (Dako, Agilent). This protocol was also used for all stainings depicted in our tumor and normal tissue galleries.
Overall, 86 of 152 tumor categories showed detectable TRPS1 staining with 36 tumor categories showing at least one strongly positive case. The TRPS1 positivity rate was highest in various types of breast cancers (51.4%-100%), soft tissue tumors (up to 100%), salivary gland tumors (up to 46.2%), squamous cell carcinomas of various sites of origin (up to 34.7%), and in diverse gynecological cancers (up to 40.0%). The distribution of positive staining results is shown in an “organ-systematic” (Figure 1) and in a “ranking order” figure (Figure 2) below (images based on data from Lennartz et al). Data on associations with histopathological and clinical parameters are also summarized below (Table 3; based on data described by Lennartz et al).
Authors conclusions on diagnostic utility of TRPS1 IHC with respect to the distinction of different tumor entities (Lennartz et al):
- TRPS1 immunostaining is a sensitive but not a fully specific marker for tumors derived from the breast.
- TRPS1 positivity is a marker for synovial sarcoma (positive in 80-100%).
- The combined analysis of TRPS1 and GATA3 is useful: A combined GATA3 and TRPS1 positivity was almost exclusively seen in neoplasms of the breast and the salivary glands. Only 2.5% of 1,159 GATA3/TRPS1 dual positive tumors were of non-breast and non-salivary gland origin in the Lennartz study.
Authors conclusions on the prognostic role of TRPS1 immunostaining results (Lennartz et al):
- Low TRPS1 expression was linked to high grade (p=0.0547), advanced pT stage (p<0.0001), nodal metastasis (p=0.0571), loss of estrogen receptor (ER) expression (p<0.0001), loss of progesterone receptor (PR) expression (p<0.0001), and triple-negative status (p<0.0001) in breast cancer of no special type.
- Low TRPS1 expression was linked to high pT (p<0.0001), nodal metastasis (p<0.0001), and high grade (p<0.0001) in a combined analysis of squamous cell carcinomas from 11 different sites.
- The rate of TRPS1 positivity increased from Gleason grade 3+3 (0%) to Gleason grade 5+5 (3.8%) and recurrent carcinomas under therapy (7.6%; p=0.0164) in prostate cancer.
The authors specifically highlighted a diagnostic utility of combining TRPS1 IHC with GATA3 immunostaining. They were able to provide such data because the same set of tumors had earlier been analyzed for GATA3 by using our MSVA-450M antibody (GATA3 Expression in Human Tumors: A Tissue Microarray Study on 16,557 Tumors, Reiswich et al. PMID: 36649695)
The figures below show the combined data on TRPS1 and GATA3 in an organ systematic (Figure 3) and a “ranking order” figure (Figure 4). Moreover, ranking orders are given for TRPS1 positive tumors within GATA3 positive (Figure 5) and GATA3 negative (Figure 6) subgroups.
Data from the publication: “TRPS1 is a highly sensitive marker for breast cancer: A tissue microarray study evaluating more than 19,000 tumors from 152 different tumor entities” Published by Lennartz et al in Am J Surg Pathol. 2024 Apr 22. PMID: 38647255 summarized in own graphics.
Figure 1. TRPS1 staining in cancer (“organ-systematic”; according to Lennartz et al.)
Figure 2. TRPS1 staining in cancer (“ranking list”; according to Lennartz et al.)
Figure 3. TRPS1 negative / GATA3 positive staining in cancer (“organ-systematic”; according to Lennartz et al.)
Figure 4. TRPS1 negative / GATA3 positive staining in cancer (“ranking order”; according to Lennartz et al.)
Figure 5. TRPS1 positive / GATA3 negative staining in cancer (“organ-systematic”; according to Lennartz et al.)
Figure 6. TRPS1 positive / GATA3 negative staining in cancer (“ranking order”; according to Lennartz et al.)
Protocol Recommendations
IHC users have different preferences on how the stains should look like. Some prefer high staining intensity of the target stain and even accept some background. Others favor absolute specificity and lighter target stains. Factors that invariably lead to more intense staining include higher concentration of the antibody and visualization tools, longer incubation time, higher temperature during incubation, higher temperature and longer duration of the heat induced epitope retrieval (slide pretreatment). The impact of the pH during slide pretreatment has variable effects and depends on the antibody and the target protein.
All images and data shown here and in our image galleries are obtained by the manual protocol described below. Other protocols resulting in equivalent staining are described as well.
Manual protocol
Freshly cut sections should be used (less than 10 days between cutting and staining). Heat-induced antigen retrieval for 5 minutes in an autoclave at 121°C in pH 7,8 Target Retrieval Solution buffer. Apply MSVA-512R at a dilution of 1:150 at 37°C for 60 minutes. Visualization of bound antibody by the EnVision Kit (Dako, Agilent) according to the manufacturer’s directions.
Potential Research Applications
- The role of TRPS1 in the biology of breast cancer and other tumors needs to be investigated.
- The role of TRPS1 in epithelial mesenchymal transition is under investigation.
- The diagnostic utility of TRPS1 immunohistochemistry awaits further clarification.
Evidence for Antibody Specificity in IHC
There are two ways how the specificity of antibodies can be documented for immunohistochemistry on formalin fixed tissues. These are: 1. Comparison with a second independent method for target expression measurement across a large number of different tissue types (orthogonal strategy), and 2. Comparison with one or several independent antibodies for the same target and showing that all positive staining results are also seen with other antibodies for the same target (independent antibody strategy).
Orthogonal validation: For the antibody MSVA-512R, a comparison with RNA data with data from three independent RNA screening studies, including the Human Protein Atlas (HPA) RNA-seq tissue dataset, the FANTOM5 project, and the Genotype-Tissue Expression (GTEx) project, which are all summarized in the Human Protein Atlas (Tissue expression TRPS1) is consistent with a specific staining as immunostaining by MSVA-512R. At least TRPS1 immunostaining was strongest in the breast (the organ with highest RNA expression). Given the abundance of RNA expression across tissues, orthogonal validation is of little help for specificity assessment of TRPS1 antibodies.
Comparison of antibodies: Full specificity of our antibody MSVA-512R is strongly supported, however, by confirmation of all cell types stained by MSVA-512R by a comparison with a second, commercially available, independent antibody (termed “validation antibody”). Comparative stainings on consecutive tissue sections are documented below.
Independence of the two antibodies is documented by cytoplasmic staining of basal cells in squamous epithelium which was only seen by the validation antibody but not by MSVA-512R.